Benchmarks and Metrics#
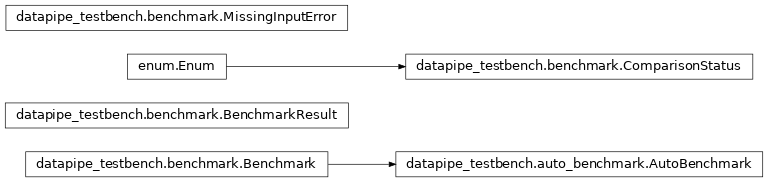
Benchmarks#
Defines what is a Benchmark.
- class Benchmark[source]#
Bases:
objectDefines what a “scientist” should define for a given benchmark.
A benchmark is defined by two functions:
generate_metrics(), which will be used to transform event data into one or moreMetricstored in aMetricsStorecorresponding to a single set of processed events.compare_to_reference(), which compares one or more MetricsStores to a reference MetricsStore. This function is run after there are multiple MetricsStores available that have been processed withgenerate_metrics.
- check_input_dataset(input_dataset: InputDataset)[source]#
Ensure required inputs exist, or raise MissingInputError.
- abstractmethod compare_to_reference(metric_store_list: list[MetricsStore], result_store: ResultStore) dict | None[source]#
Perform the comparison study on metrics associated with a set of experiments.
This function takes a list of MetricStores that have been previously filled by
datapipe_testbench.benchmark.Benchmark.generate_metrics(), generates plots and comparison results, and writes the output to a ResultsStore. If there are more than one MetricStore to compare, the first one is considered the _reference_.- Parameters:
- metric_store_list: list[MetricsStore]
The list of MetricStores to compare, the first of which is the _reference_ .
- result_store: ResultStore
WHere the results of the comparison study are stored.
- abstractmethod generate_metrics(metric_store: MetricsStore) dict | None[source]#
Produce metrics for this benchmark later comparison.
Called once per benchmark per
MetricsStore, i.e. for one set of input events, defines how to transform the event into to a metrics stored in aMetricsStorethat can later be compared.- Parameters:
- metric_store: MetricsStore
Where to store the metrics generated by this
Benchmark. It must be initialized with andatapipe_testbench.inputdataset.InputDatasetcontaining the inputs to use for the transformation into metrics.
- abstractmethod make_report(result_store: ResultStore)[source]#
Make a report for the benchmark using the results of the comparisons.
The result_store must already contain the outputs of
compare_to_reference().- Parameters:
- result_storeResultStore
Contains the input comparisons and is where the report will be stored.
- missing_outputs(metric_store: MetricsStore)[source]#
Return list of missing outputs.
- property name#
Return friendly name of this benchmark.
- class BenchmarkResult(benchmark_name: str, reference_dataset_name: str, test_dataset_name: str, status: ComparisonStatus, comment: str, plots: list[FigureBase] | None)[source]#
Bases:
objectOutput of a benchmark is one or more of these.
- class ComparisonStatus(*values)[source]#
-
Status of a benchmark check.
- FAILED = '3'#
benchmark failed
- OTHER = '4'#
other failure, or couldn’t complete benchmark
- PASSED = '1'#
benchmark passed
- WARNING = '2'#
benchmark passed, but only barely, should check
- exception MissingInputError[source]#
Bases:
RuntimeErrorRaised when required inputs are missing.
Metrics#
Ndhistogram metric and datamodel location definition.
- class Metric(axis_list: list[AxesMixin], unit: Unit | str = '', label: str | None = None, output_store: Storable | None = None, other_attributes: dict | None = None, data=None, hist: Hist | None = None)[source]#
Bases:
StorableND-Histogram that can be serialized to ASDF.
Class stores ND-histogram based on scikit-hep Hist package to create bins by accumulation.
This initializer should not be overridden in subclasses! If you need a specialized constructor, define a @classmethod that does what is needed and calls this generic one to return a properly constructed Metric. Otherwise serialization will not work.
- Parameters:
- axis_listlist[axis.AxesMixin]
List of hist.axis objects. Each can define a metadata keyword (str) that will be used as the. If the axis has a unit, set it in the “unit” key in the axis.metadata dict
- labelstr | None
if none, an automatic label will be generated.
- unit: u.Unit | None:
Unit of the _data_ stored in the metric. Note that you should set _axis_ units by passing them as metadata to the axes in the axis_list, i.e.
RegularAxis(..., metadata=dict(unit="deg"))- output_store: ResultStore | None
If specified, the name of this metric will be updated to reflect the output name. This is used e.g. by AutoBenchmark for nicer bookkeeping and to avoid name collisions.
- other_attributes: dict[str,Any] | None
Other attributes of this metric that should be stored along with it.
- data: np.array | None
If specified, will set the internal histogram to this value.. Must be the same shape as the axes specify.
- hist: Hist | None
If set, use this for the internal histogram rather than constructing one. Must have the same axis definition as specified .
- property axes#
Wrapper to transfer to _storage.
- abstractmethod compare(others: list)[source]#
Compare several instances of the class.
Uses
selfas reference when comparing withothers. Plots the spectra on top of each other, and calculates the wasserstein distance.- Parameters:
- otherslist[Metric]
List containing other instances of this class.
- Returns:
- ComparisonResult | None
Results of comparison of this reference to the others list.
- compatible(other)[source]#
Compare objects and ensure number and name of axis match.
- Parameters:
- other
- Metric other
Other instance to compare self to.
- Returns:
- True if the objects are compatible.
- Raises:
- IndexError.
- get_identifier()[source]#
Auto generation of a unique identifier for the metric.
Use axis order and name to return a tuple that need to be unique and allow identification of the current metric throughout multiple metric stores.
- normalize_to_probability(axis_index: int = 1, threshold_frac=0.005, fill_value=nan) Self[source]#
Normalize the data such that the integral over axis
axis_index1.0.- Parameters:
- arrnp.ndarray
The input array.
- axis_indexint
The axis along which to normalize.
- threshold_frac: float
if the integral along the axis is below this fraction of the total integral, mask off this row as having too low stats to plot. This prevents low-stats values form saturating the color scale
- fill_value:
value to replace low-stats entries with
- Returns:
- np.ndarray
A new Metric normalized along the specified axis.
- plot_compare_1d(others: Self | list[Self] = None, fig: None | FigureBase = None, legend=True, show_xlabel=True, layout: str | None = 'constrained', sharex: Axes | None = None, sharey: Axes | None = None, sharey_diff: Axes | None = None, **kwargs) dict[str, Axes][source]#
Generate a subfigure with comparisons and residuals.
The residuals are the relative differences, \(\Delta_\mathrm{rel} \equiv \frac{(v_i - v_\mathrm{ref})}{v_\mathrm{ref}}\)
- Parameters:
- others: Metric | list[Metric] | None
List of metrics to compare to reference
- fig: matplotlib.Figure | None
Figure or SubFigure to add the axes to, or none to use current.
- legend: bool
Show legend on plot
- layout: str | None
If fig not passed, in, use this maptlotlib layout engine.
- sharex: Axes | None
If specified, share all x-axes with this axis.
- sharey: Axes | None
If specified share the value’s y-axis with this axis
- sharey_diff: Axes | None
If specified share the diff’s y-axis with this axis.
- **kwargs:
Any other args are passed to the plot() functions.
- Returns:
- dict[matplotlib.Axes]:
“val”: value_axis, “diff”: relative_difference_axis
- abstractmethod classmethod setup(*args, **kwargs) Self[source]#
Create a new Metric with specialized parameters.
Overridden by subclasses to properly setup the Metric the first time with user-specified and metric-specific parameters. This should be called instead of the default constructor to set up the Metric correctly. Afterward, it will be loaded from a file using the default Metric constructor. This is because all Metrics must share the same
__init__method for serialization to work.

